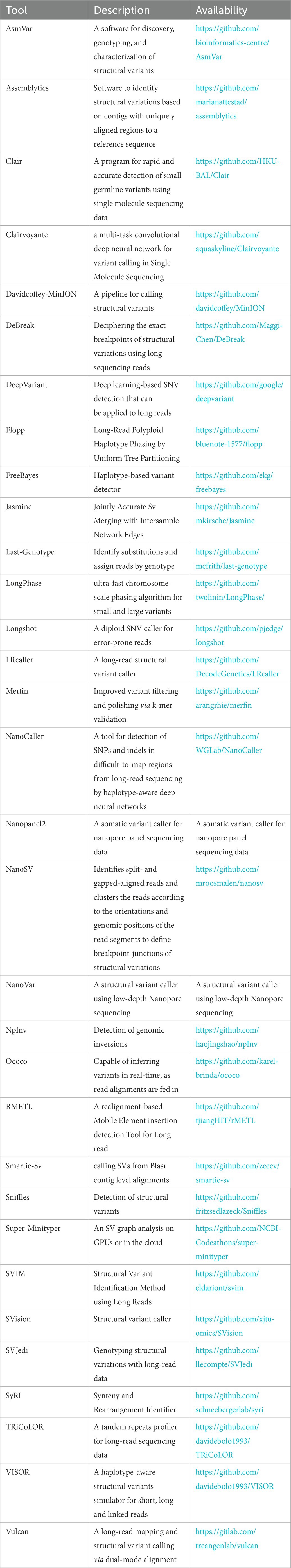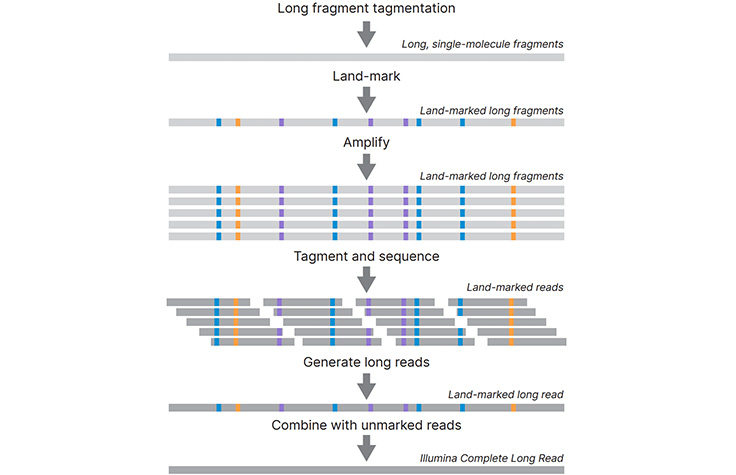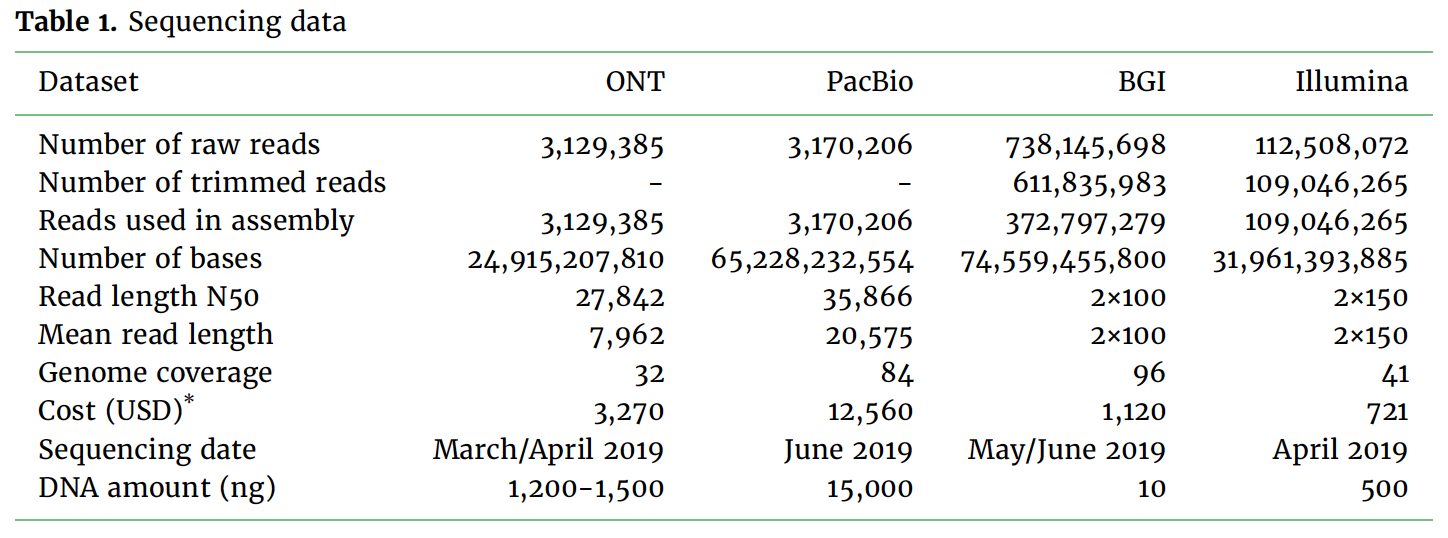
Benchmarking of long read sequencing technologies and genome assembly tools - Genome Innovation Hub - University of Queensland

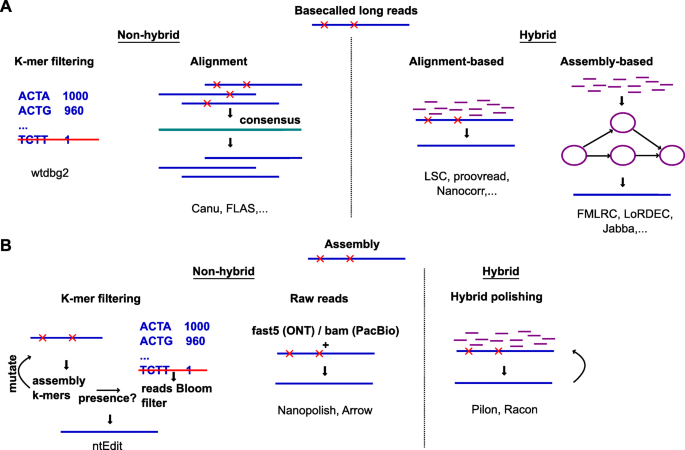
Overview of long-read analysis tools and pipelines. a Release of tools... | Download Scientific Diagram
Linear time complexity de novo long read genome assembly with GoldRush,Nature Communications - X-MOL

PacBio on X: "We asked @sedlazeck to share his motivations for developing new algorithms for structural variant detection using long-read sequencing. Check out the Q&A on our blog: https://t.co/mytbGDl6RL https://t.co/awAhSRO37S" / X
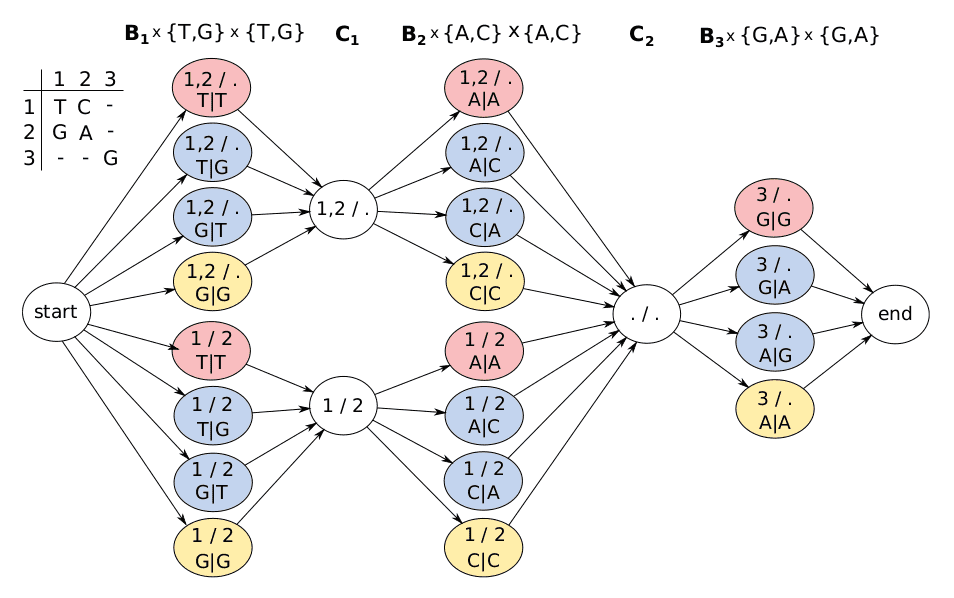
Haplotype-aware genotyping from long reads with Trevor Pesout — the bioinformatics chat, a podcast about bioinformatics
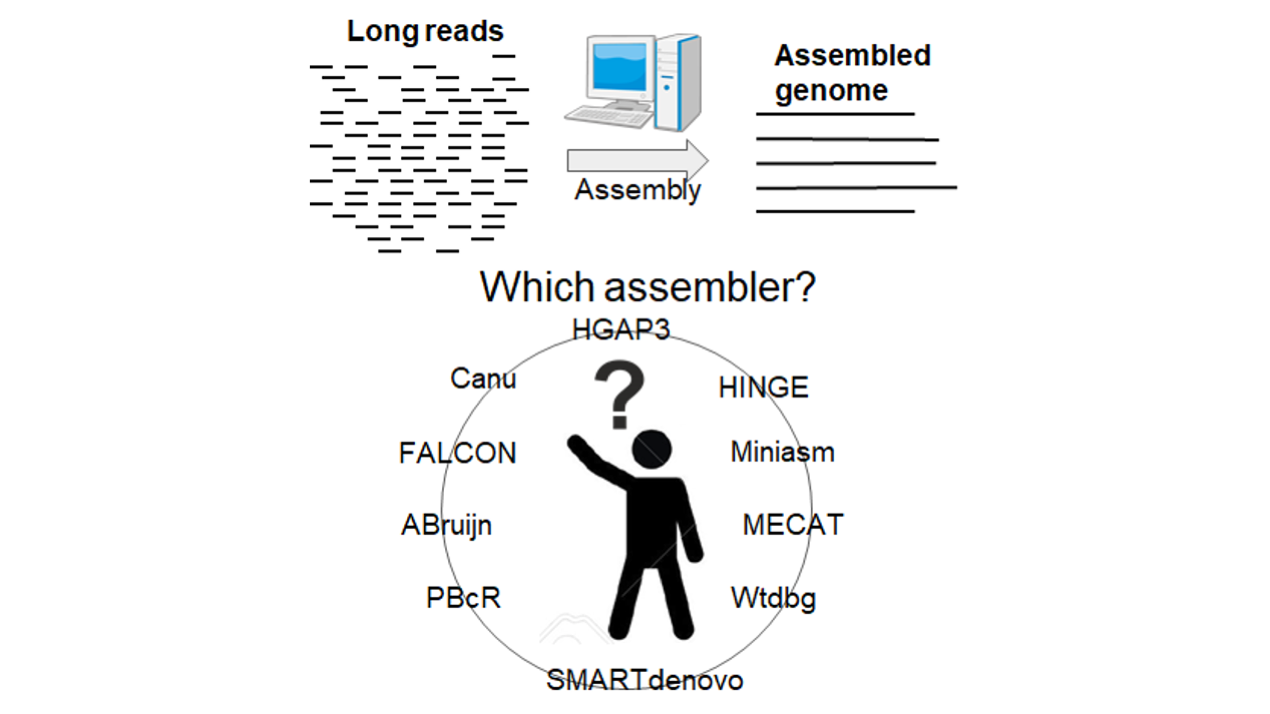
Comprehensive evaluation of non-hybrid genome assembly tools for third-generation PacBio long-read sequence data | 慶應義塾大学 グローバルリサーチインスティテュート

Hidden in Plain Sight — A Bioinformatics Journey into Structural Variation Calling in the Long Read Sequencing Era | by Liz T | PacBio | Medium
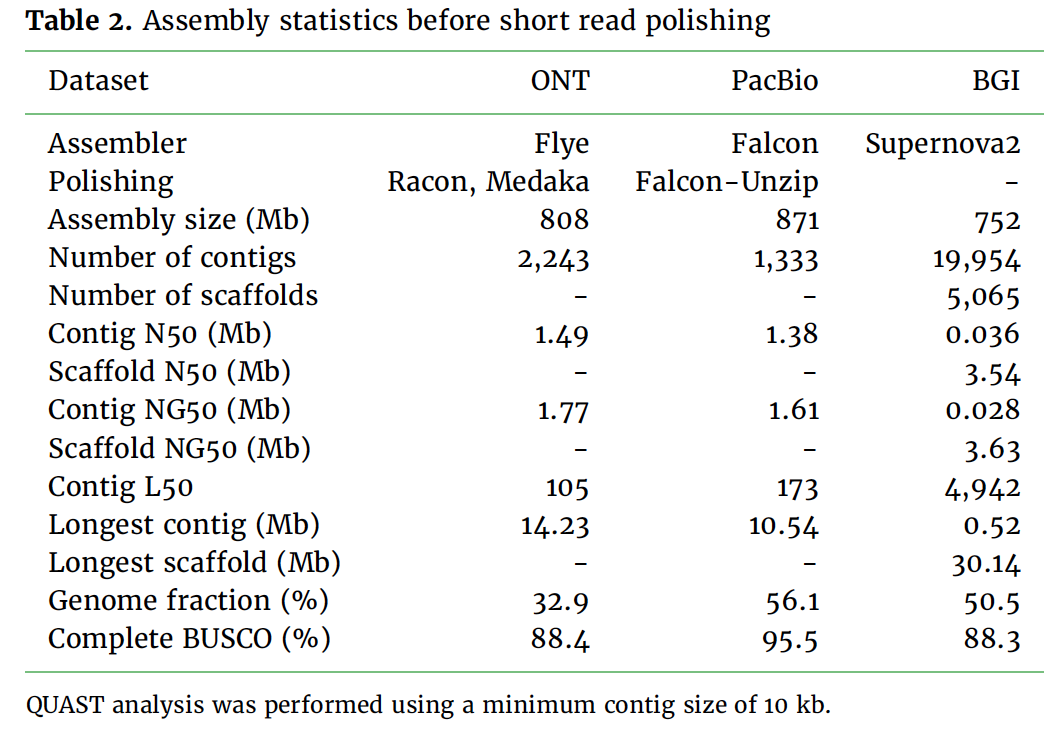
Benchmarking of long read sequencing technologies and genome assembly tools - Genome Innovation Hub - University of Queensland








![PDF] NanoGalaxy: Nanopore long-read sequencing data analysis in Galaxy | Semantic Scholar PDF] NanoGalaxy: Nanopore long-read sequencing data analysis in Galaxy | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c29adb742dd67908f2cd858e1a8bad85e50a7cfb/11-Table1-1.png)